SSM Procedure
Example 33.13 Bivariate Model: Sales of Mink and Muskrat Furs
(View the complete code for this example.)
This example considers a bivariate time series of logarithms of the annual sales of mink and muskrat furs by the Hudson’s Bay Company for the years 1850–1911. These data have been analyzed previously by many authors, including Chan and Wallis (1978); Harvey (1989); Reinsel (1997). There is known to be a predator-prey relationship between the mink and muskrat species: minks are principal predators of muskrats. Previous analyses for these data generally conclude the following:
An increase in the muskrat population is followed by an increase in the mink population a year later, and an increase in the mink population is followed by a decrease in the muskrat population a year later.
Because muskrats are not the only item in the mink diet and because both mink and muskrat populations are affected by many other factors, the model must include additional terms to explain the year-to-year variation.
The analysis in this example, which loosely follows the discussion in Harvey (1989, chap. 8, sec. 8), also leads to similar conclusions. It begins by taking Harvey’s model 8.8.8 (a and b), with autoregressive order one, as the starting model—that is, it assumes that the bivariate (mink, muskrat) process satisfies the following relationship:
This model postulates that can be expressed as a sum of three terms:
, a bivariate trend that is modeled as a random walk with drift
;
, an AR(1) correction; and
, a bivariate Gaussian white noise disturbance. It is assumed that the AR coefficient matrix
is stable (that is, its eigenvalues are less than 1 in magnitude) and that the bivariate disturbances
(white noise associated with
) and
are mutually independent.
The following statements show how you can specify this model in the SSM procedure:
proc ssm data=furs plots=residual;
/* Specify the ID variable */
id year interval=year;
/* Define parameters */
parms rho1 rho2/ lower=-0.9999 upper=0.9999;
parms msd1 msd2 esd1 esd2 / lower=1.e-6;
/* Specify the terms with lagged response variables */
deplag LagsForMink(LogMink) LogMink(lags=1) LogMusk(lags=1);
deplag LagsForMusk(LogMusk) LogMink(lags=1) LogMusk(lags=1);
/* Specify the bivariate trend */
array rwQ{2,2};
rwQ[1,1] = msd1*msd1; rwQ[1,2] = msd1*msd2*rho1;
rwQ[2,1] = rwQ[1,2]; rwQ[2,2]=msd2*msd2;
state alpha(2) type=RW W(I) cov(g)=(rwQ);
comp minkLevel = alpha[1];
comp muskLevel = alpha[2];
/* Specify the bivariate white noise */
array wnQ{2,2};
wnQ[1,1] = esd1*esd1; wnQ[1,2] = esd1*esd2*rho2;
wnQ[2,1] = wnQ[1,2]; wnQ[2,2]=esd2*esd2;
state error(2) type=WN cov(g)=(wnQ);
comp minkWn = error[1];
comp muskWn = error[2];
/* Specify the observation equation */
model LogMink = LagsForMink minkLevel minkWn;
model LogMusk = LagsForMusk muskLevel muskWn;
/* Specify an output data set to store component estimates */
output out=salesFor press;
run;
The different parts of the program are explained as follows:
The PARMS statements define parameters that are used to form the elements of
(the covariance of
, the disturbance term in the bivariate level equation) and
(the covariance of
, which is the bivariate white noise).
is parameterized as (
msd1*msd1 msd1*msd2*rho1; msd1*msd2*rho1 msd2*msd2).is similarly parameterized by using
esd1, esd2, andrho2. In addition to ensuring thatand
are positive semidefinite, it turns out that this parameterization leads to an interpretable model at the end.
The DEPLAG statements help define the terms that are associated with
.
The remaining statements are self-explanatory.
Output 33.13.1 shows the estimate of the drift vector in the equation of
(
).
Output 33.13.1: Estimate of the Drift Vector
| Estimate of the State Equation Regression Vector | |||||
|---|---|---|---|---|---|
| State | Element Index | Estimate | Standard Error |
t Value | Pr > |t| |
| alpha | 1 | -0.000817 | 0.0323 | -0.03 | 0.9798 |
| alpha | 2 | 0.005953 | 0.0258 | 0.23 | 0.8175 |
Clearly, both elements of are statistically insignificant, and the
equation can be simplified as
. Next, Output 33.13.2 shows the estimates of the elements of
, and
, and Output 33.13.3 shows the estimates of the lag coefficients.
Output 33.13.2: Estimates of , and
| Estimates of Named Parameters | |||
|---|---|---|---|
| Parameter | Estimate | Standard Error |
t Value |
| rho1 | 0.8310 | 0.1377 | 6.03 |
| rho2 | -0.9999 | 1.6555 | -0.60 |
| msd1 | 0.2500 | 0.0354 | 7.06 |
| msd2 | 0.1991 | 0.0592 | 3.36 |
| esd1 | 0.0662 | 0.0597 | 1.11 |
| esd2 | 0.1344 | 0.0527 | 2.55 |
Output 33.13.3: Estimates of Lag Coefficients (Elements of )
| Model Parameter Estimates | |||||
|---|---|---|---|---|---|
| Component | Type | Parameter | Estimate | Standard Error |
t Value |
| LagsForMink | Lag Coefficient Of LogMink | Lag[1] | -0.0011 | 0.173 | -0.01 |
| LagsForMink | Lag Coefficient Of LogMusk | Lag[1] | 0.3349 | 0.137 | 2.45 |
| LagsForMusk | Lag Coefficient Of LogMink | Lag[1] | -0.9905 | 0.142 | -6.98 |
| LagsForMusk | Lag Coefficient Of LogMusk | Lag[1] | 0.6570 | 0.121 | 5.44 |
The main points of the output can be summarized as follows:
phi11, the first element of, which relates the current value of
LogMinkwith its lagged value, is statistically insignificant. That is, laggedLogMinkterm could be dropped from the model equation forLogMink.rho2, the correlation coefficient between the elements of—the bivariate noise vector in the equation
—is very near its lower boundary of –1 (in such cases the standard error of the parameter estimate is not reliable). This implies that the two elements of
are perfectly negatively correlated.
Taken together, these observations suggest the reduced model
where and
is parameterized as (
esd1*esd1 -esd1*esd2; -esd1*esd2 esd2*esd2). The program that produces the reduced model is a simple modification of the preceding program and is not shown.
Output 33.13.4: Estimates of , and
(Reduced Model)
| Estimates of Named Parameters | |||
|---|---|---|---|
| Parameter | Estimate | Standard Error |
t Value |
| rho1 | 0.8526 | 0.0978 | 8.71 |
| msd1 | 0.2472 | 0.0282 | 8.78 |
| msd2 | 0.1955 | 0.0365 | 5.36 |
| esd1 | 0.0679 | 0.0385 | 1.76 |
| esd2 | 0.1372 | 0.0328 | 4.18 |
Output 33.13.5: Estimates of Lag Coefficients (elements of ) (Reduced Model)
| Model Parameter Estimates | |||||
|---|---|---|---|---|---|
| Component | Type | Parameter | Estimate | Standard Error |
t Value |
| LagsForMink | Lag Coefficient Of LogMusk | Lag[1] | 0.330 | 0.0986 | 3.35 |
| LagsForMusk | Lag Coefficient Of LogMink | Lag[1] | -0.997 | 0.1168 | -8.54 |
| LagsForMusk | Lag Coefficient Of LogMusk | Lag[1] | 0.668 | 0.1003 | 6.66 |
The tables in Output 33.13.4 and Output 33.13.5 show the new parameter estimates. By examining the parameter estimates, you can easily see that this model supports the general conclusions mentioned at the start of this example. In particular, note the following:
implies that this year’s mink abundance is positively correlated with last year’s muskrat abundance.
and
imply that this year’s muskrat abundance is negatively correlated with last year’s mink abundance and positively correlated with last year’s muskrat abundance.
Even though the parameters were not restricted to ensure stability, the estimated
turns out to be stable with a pair of complex eigenvalues,
, and a modulus of 0.570 (these calculations are done separately by using the IML procedure).
The fact that elements of
are perfectly negatively correlated further supports the predator-prey relationship.
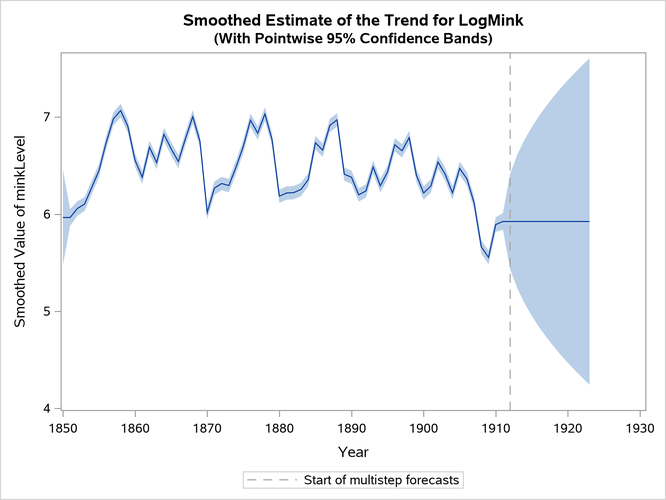
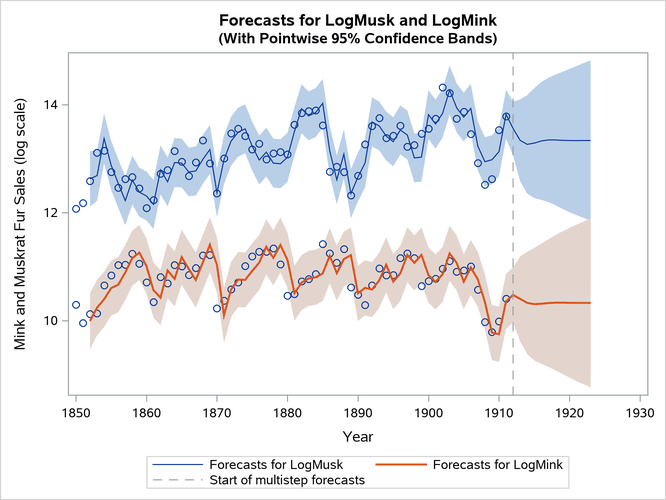
Finally, Output 33.13.6 shows the plots of one-step-ahead and post-sample forecasts for LogMink and LogMusk, and Output 33.13.7 shows the plot of the smoothed (full-sample) estimate of the first element of : LogMink Trend.
Output 33.13.6: Forecasts for Mink and Muskrat Fur Sales in Logarithms
Output 33.13.7: Smoothed Estimate of LogMink Trend